When it comes to death rates for invasive European rabbits (Oryctolagus cuniculus) in Australia, it appears that 1 + 1 = 2.1.

Tagged European rabbit kitten infected with myxoma virus, but that died from rabbit haemorrhagic virus disease (RHDV). Photo by David Peacock, Biosecurity South Australia.
“Canberra, we have a problem” — Sure, it’s an old problem and much less of one than it used to be back in the 1950s, but invasive rabbits are nonetheless an ecological, conservation, and financial catastrophe across Australia.
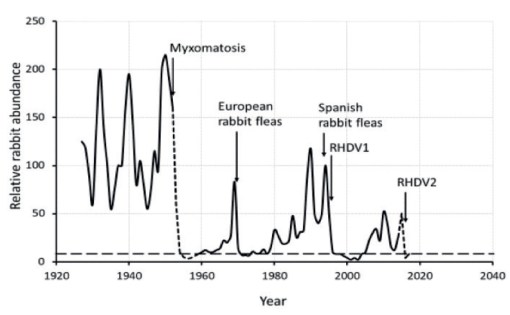
Semi-schematic diagram, redrawn using data from Saunders and others and extended to include the recent spread of RHDV2, showing changes in rabbit abundance in relation to the introduction of biological control agents into north-eastern South Australia. Dotted lines indicate uncertainty due to lack of continuous annual data. The broken line indicates a level of about 0.5 rabbits ha-1, below which rabbits must be held to ensure recovery of native pastures and shrubs (from B. Cooke 2018 Vet Rec doi:10.1136/vr.k2105)
Rabbits used to reach plague numbers in much of agricultural and outback Australia, but the introduction and clever manipulation of two rather effective rabbit-specific viruses and insect vectors — first, myxoma virus in 1950, European rabbit fleas in the 1960s to help spread the virus, then Spanish rabbit fleas in the 1990s to increase spread into arid areas, and then rabbit haemorrhagic disease virus (RHDV) in 1995 — have been effective in dropping rabbit abundances by an estimated 75-80% in South Australia alone since the 1950s.




